The F-Distribution¶
The F-Distribution is used in statistics to test whether the difference between the variance of two samples is statistically significant. For example, the standard deviation in how close a marksman hits a target along the x-coordinate may vary between two people, also known as "precision" in marksmanship lingo.
It is also notably used for Analysis of Variance (ANOVA) testing, which tests whether samples from multiple different categories are statistically different from the whole set of categories by comparing variance between different categories with the variance within categories.
The F-statistic is calculated by the ratio of the variances (or square of the standard deviations): $$F=\frac{s_1^2/d_1}{s_2^2/d_2}$$ Where these standard deviations, $s$, are sometimes denoted as "numerator" and "denominator", and $d$ is the corresponding degrees of freedom.
The "F-distribution" may refer to the probability distribution function (PDF), cumulative distribution function (CDF), or survival function (SF=1-CDF), depending on how you interpret it. These functions depends on the degrees of freedom for both samples as well as the significance level, $\alpha$ (the area of the tail in the PDF).
The F-tables usually provide the F-value corresponding to where the "right-tail" integral of the PDF or value of the SF=1-CDF is equal to $\alpha$. There may be F-tables that provide the "left-tail" integral, in which case, those provide the F-value corresponding to where the CDF=$\alpha$
from scipy.stats import f
import numpy as np
from matplotlib import pyplot as plt
dfnlist = np.arange(7,23,4)
dfdlist = np.arange(7,23,4)
plt.figure(figsize=(10,10))
q = 0.05
plt.subplot(3,2,1)
dfn = dfnlist[0]
for i in range(len(dfdlist)):
dfd = dfdlist[i]
x = np.linspace(f.ppf(q,dfn,dfd),f.ppf(1-q,dfn,dfd),1000)
pdf = f.pdf(x,dfn,dfd)
plt.plot(x,pdf,label=f'dfd={dfd}')
plt.title(f'dfn={dfn}')
plt.legend()
plt.xlabel('F')
plt.ylabel('PDF')
plt.subplot(3,2,2)
dfd = dfdlist[0]
for i in range(len(dfnlist)):
dfn = dfnlist[i]
x = np.linspace(f.ppf(q,dfn,dfd),f.ppf(1-q,dfn,dfd),1000)
pdf = f.pdf(x,dfn,dfd)
plt.plot(x,pdf,label=f'dfn={dfn}')
plt.title(f'dfd={dfd}')
plt.legend()
plt.xlabel('F')
plt.ylabel('PDF')
plt.subplot(3,2,3)
dfn = dfnlist[0]
for i in range(len(dfdlist)):
dfd = dfdlist[i]
x = np.linspace(f.ppf(q,dfn,dfd),f.ppf(1-q,dfn,dfd),1000)
pdf = f.cdf(x,dfn,dfd)
plt.plot(x,pdf,label=f'dfd={dfd}')
plt.title(f'dfn={dfn}')
plt.legend()
plt.xlabel('F')
plt.ylabel('CDF')
plt.subplot(3,2,4)
dfd = dfdlist[0]
for i in range(len(dfnlist)):
dfn = dfnlist[i]
x = np.linspace(f.ppf(q,dfn,dfd),f.ppf(1-q,dfn,dfd),1000)
pdf = f.cdf(x,dfn,dfd)
plt.plot(x,pdf,label=f'dfn={dfn}')
plt.title(f'dfd={dfd}')
plt.legend()
plt.xlabel('F')
plt.ylabel('CDF')
plt.subplot(3,2,5)
dfn = dfnlist[0]
for i in range(len(dfdlist)):
dfd = dfdlist[i]
x = np.linspace(f.ppf(q,dfn,dfd),f.ppf(1-q,dfn,dfd),1000)
pdf = f.sf(x,dfn,dfd)
plt.plot(x,pdf,label=f'dfd={dfd}')
plt.title(f'dfn={dfn}')
plt.legend()
plt.xlabel('F')
plt.ylabel('SF=1-CDF')
for i in range(len(dfnlist)):
dfd = dfnlist[i]
x = f.ppf(1-q,dfn,dfd)
plt.plot(x,f.sf(x,dfn,dfd),'k.')
plt.subplot(3,2,6)
dfd = dfdlist[0]
for i in range(len(dfnlist)):
dfn = dfnlist[i]
x = np.linspace(f.ppf(q,dfn,dfd),f.ppf(1-q,dfn,dfd),1000)
pdf = f.sf(x,dfn,dfd)
plt.plot(x,pdf,label=f'dfn={dfn}')
plt.title(f'dfd={dfd}')
plt.legend()
plt.xlabel('F')
plt.ylabel('SF=1-CDF')
plt.tight_layout()
plt.show()
F Table¶
Notice in the F-table that for dfn=df1=7, the F-table ($\alpha=0.05$) gives for the following values of dfd=df2:
$$7, 11, 15, 19$$
the values of the following (respectively):
$$3.79, 3.01, 2.71, 2.54$$
This is correctly reflected in the plot (black dots) of the survival function for dfn=7.
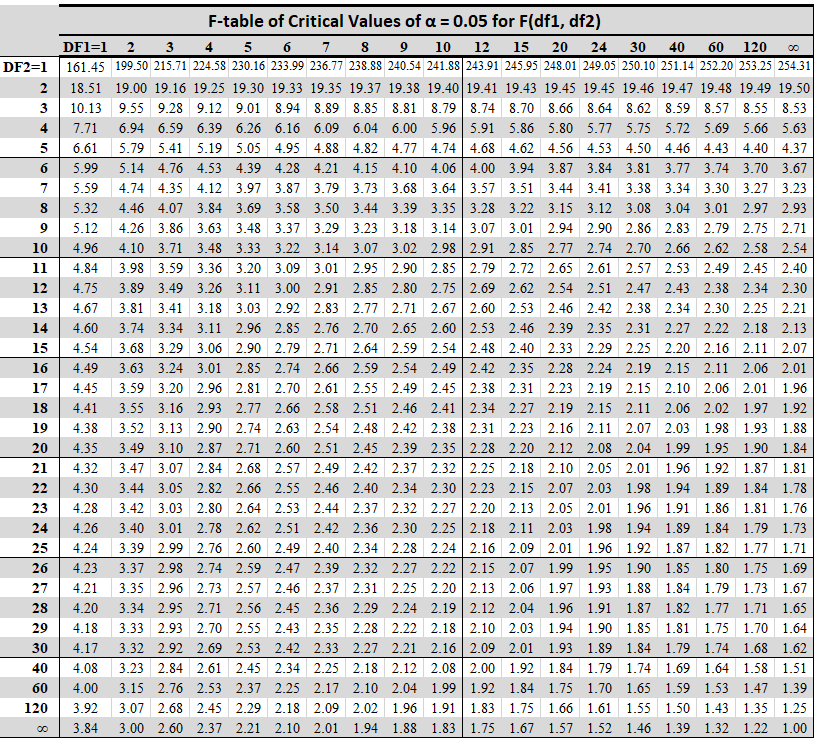 Source: https://statisticsbyjim.com/hypothesis-testing/f-table/
Source: https://statisticsbyjim.com/hypothesis-testing/f-table/